Need to survive in a tough environment? There’s an app for that
Artist’s depiction of Prochlorococcus cells in ocean water. (Image by Derek Tan, used under Creative Commons BY-NC-ND.)
For Massachusetts Institute of Technology professor Penny Chisholm, the most exciting apps will not download to your phone. Only bacteria can run them.
Chisholm studies a group of photosynthesizing bacteria that developed what she describes as their own “app store”—a set of genes shared and swapped between individual cells that enable them to thrive in a range of environmental conditions wider than what their competitors can tolerate.
The bacteria, in the genus Prochlorococcus, are the crucial first strand in the marine food web. Surveys estimate the total Prochlorococcus population amounts to more than 3 billion billion billion—that is a three followed by 27 zeroes. Those huge numbers are necessary to sustain an array of life forms, from microscopic plankton to humpback whales.
Ubiquitous, yet long unseen
Yet because of its minuscule size, Prochlorococcus was not discovered until the 1980s.
“There was nothing supposed to be that small,” Chisholm said in a presentation during CASW’s New Horizons in Science briefings for science writers Oct. 11 at MIT in Cambridge, MA, as part of ScienceWriters2015. “It’s a wonder anyone knew to look at what looked like schmutz under the microscope.”
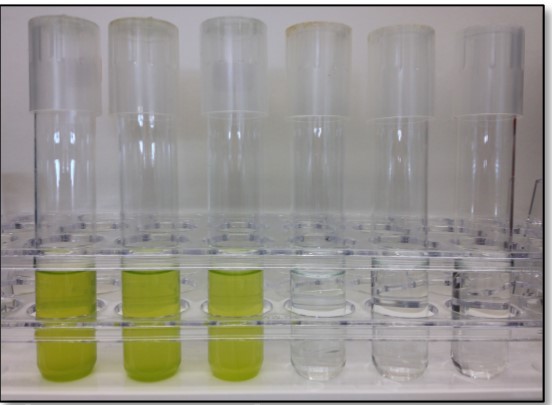

But Rob Olsen, then a postdoctoral researcher in Chisholm’s lab, kept seeing that schmutz in sea water samples. When the supposedly lifeless seawater samples were fed nutrients, they bloomed into the unmistakable green of photosynthesis in motion. When Olsen examined the samples closely under a microscope, he saw cells around one millionth of a meter in diameter—the smallest photosynthesizing organisms on the planet.
At first, research focused on the seemingly endless variation in different strains of Prochlorococcus. Some were adapted to cold water, some to warm water, some to the intense light of the ocean’s surface, some to the barely-lit gloom of deeper water. Still others were adapted to the variability in light created by the movement of currents.
But a surprise came in 2002, when the first genome sequences for Prochlorococcus became available. Each strain had around 2,000 genes, but as more and more sequences were completed, the total number of unique genes kept climbing. After more than a decade of sequencing, each new Prochlorococcus genome studied adds another 100-200 previously unknown genes to the total pool. Chisholm now estimates that the total number of unique genes possessed by Prochlorococcus might be as high as 80,000, four the size of the human genome.
A tiny operating system that challenges our view of evolutionary adaptation
Chisholm has determined that each Prochlorococcus cell has an extremely stripped-down “operating system” of around a thousand genes required to perform the basic cell operations, with the rest comprising its complement of those “apps”—genes that can give advantages or additional functions depending on the environment in which the cell is living. App genes can help secure nutrients, protect against bright light or improve survival in low temperatures, although the function of most of them is currently unknown.
This system appears to be unique to Prochlorococcus and is key to their ability to thrive everywhere from the balmy equatorial waters of the Indian Ocean to the tempest-strewn Tasman Sea. Different suites of “apps” optimize the cells to live in different environments. Cold-tolerance genes and chlorophyll boosters aid populations living in deep water and at high latitudes. Nutrient-acquisition genes go a long way in the iron-poor open waters of the Pacific.
Chisholm says Prochlorococcus changed her perspective on evolution. Previously, researchers assumed that organisms carried all their genes with them and that natural selection focused on different gene variants, like those that affect beak size in finches or eye color in humans. Prochlorococcus upends this assumption. Its compact individual genome allows it to function efficiently while still having access to a range of adaptations that dwarfs our own and which would be impossible for any single cell to maintain. Furthermore, adaptations don’t only spread vertically from one generation to the next, but horizontally between different groups. For Chisholm, understanding Prochlorococcus requires integrating research at the cellular, local, regional, and global scales.
“I call it a masterpiece because it blows my mind when I look at it,” Chisholm said. “Life has the iPhone beat.”